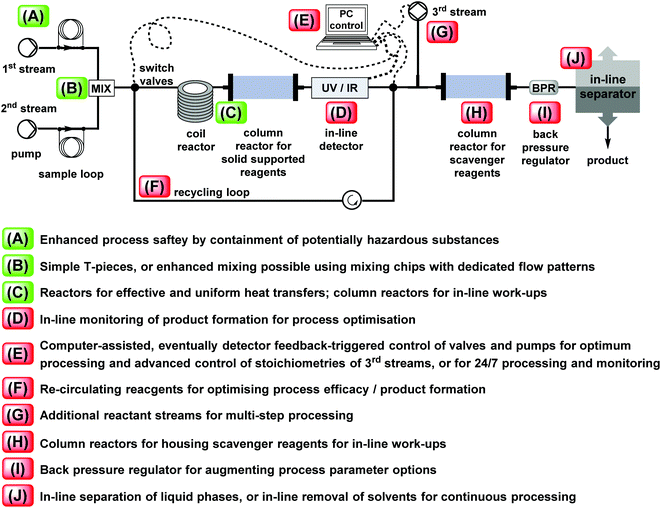
Flow synthesis approaches to privileged scaffolds – recent routes reviewed for green and sustainable aspects - Green Chemistry (RSC Publishing) DOI:10.1039/D0GC03883K

PDF) Deletion of Ubiquitin Fold Modifier Protein Ufm1 Processing Peptidase Ufsp in L. donovani Abolishes Ufm1 Processing and Alters Pathogenesis
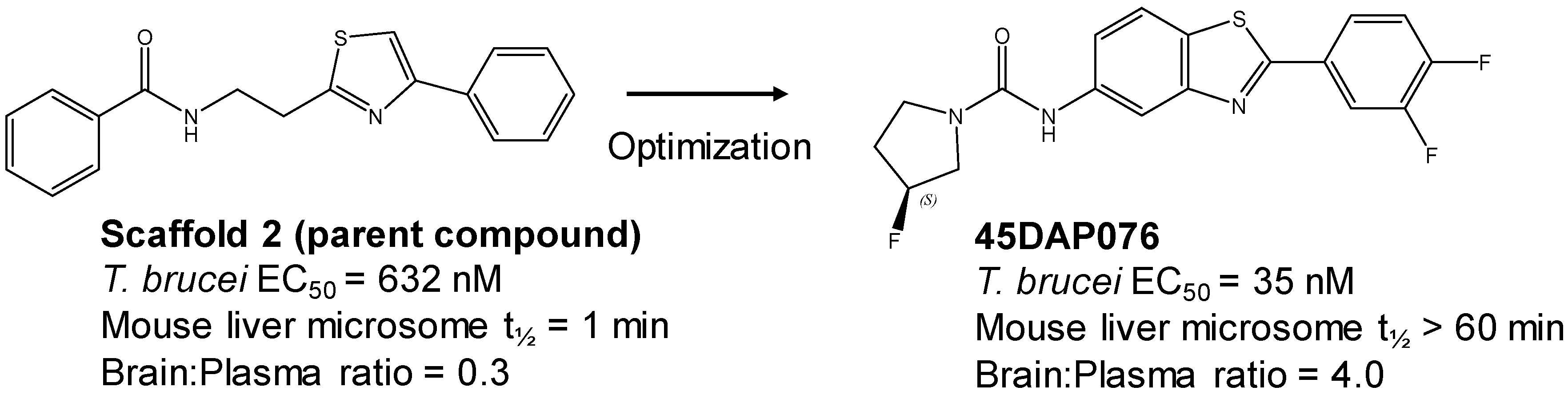
TropicalMed | Free Full-Text | Phenotypic Drug Discovery for Human African Trypanosomiasis: A Powerful Approach | HTML
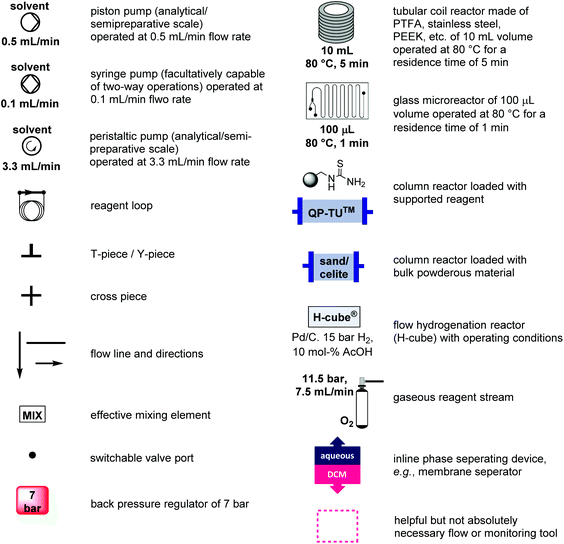
Flow synthesis approaches to privileged scaffolds – recent routes reviewed for green and sustainable aspects - Green Chemistry (RSC Publishing) DOI:10.1039/D0GC03883K

Göteborg International Biennial for Contemporary Art 2019 Guide English by Röda Sten Konsthall - Issuu

(PDF) Whole proteome analysis of human tankyrase knockout cells reveals targets of tankyrase-mediated degradation
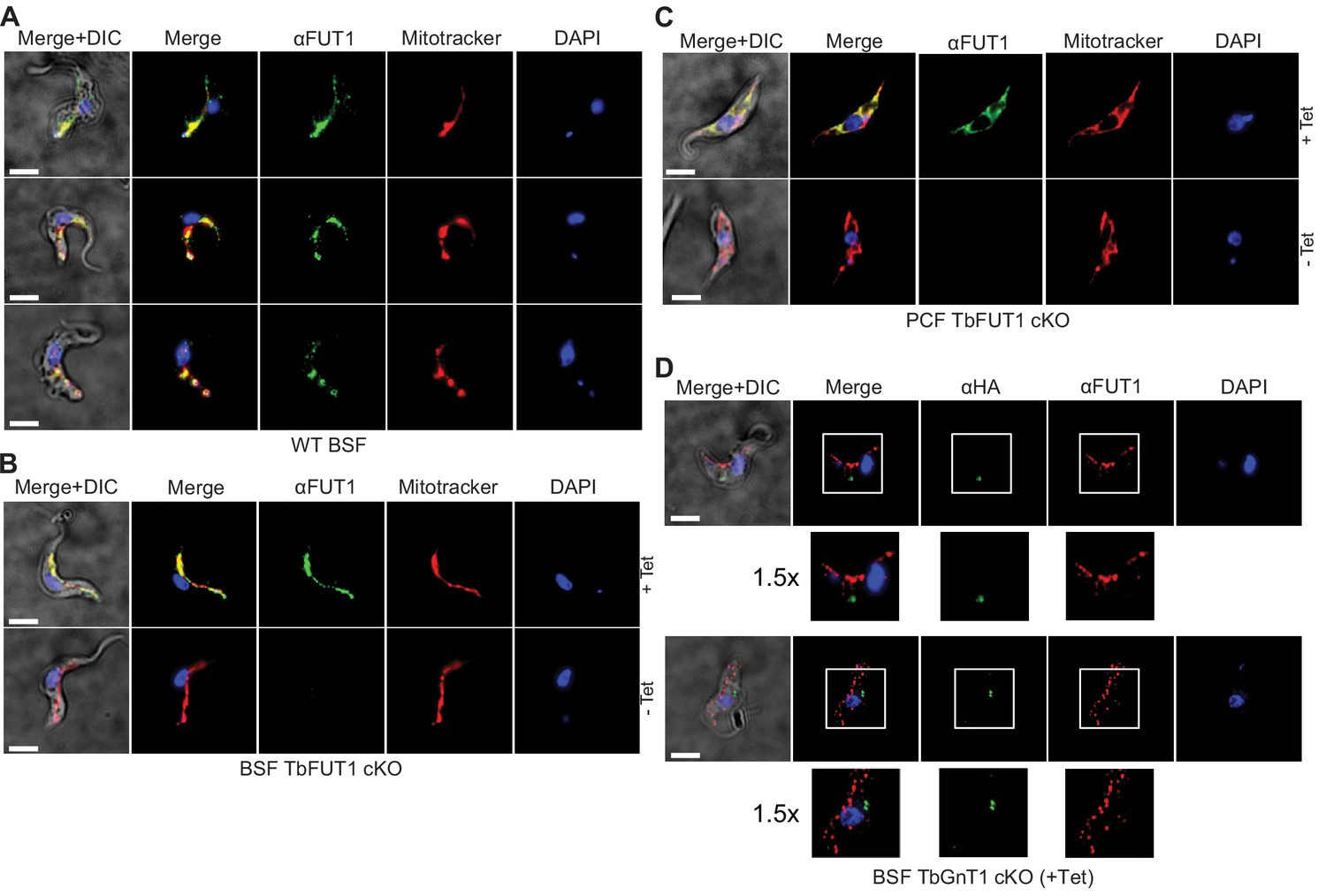
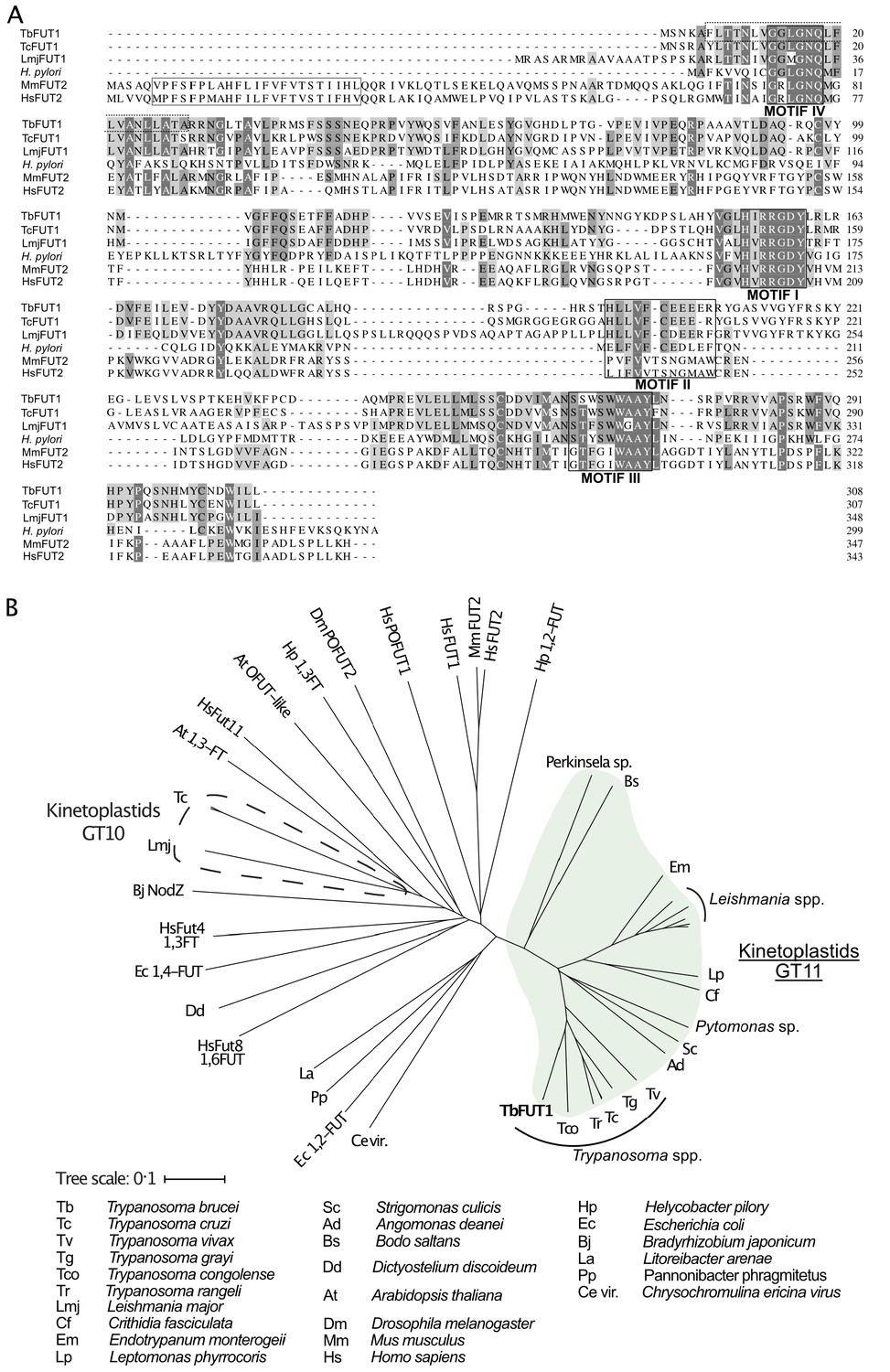
An essential, kinetoplastid-specific GDP-Fuc: β-D-Gal α-1,2-fucosyltransferase is located in the mitochondrion of Trypanosoma brucei | eLife



:no_upscale()/cdn.vox-cdn.com/uploads/chorus_asset/file/23039076/Screen_Shot_2021_11_23_at_17.16.11.png?w=715&ssl=1)